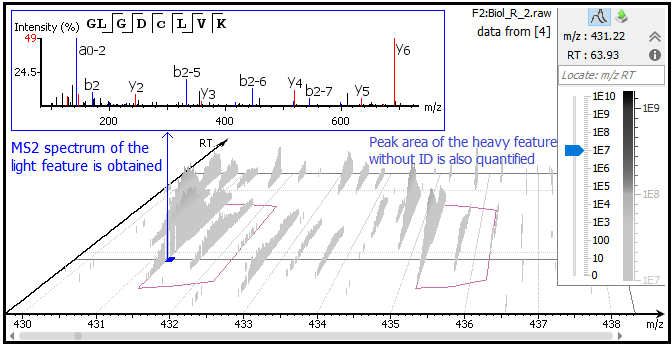
Use of stable isotope labeling by amino acids in cell culture as a spike-in standard in quantitative proteomics | Nature Protocols
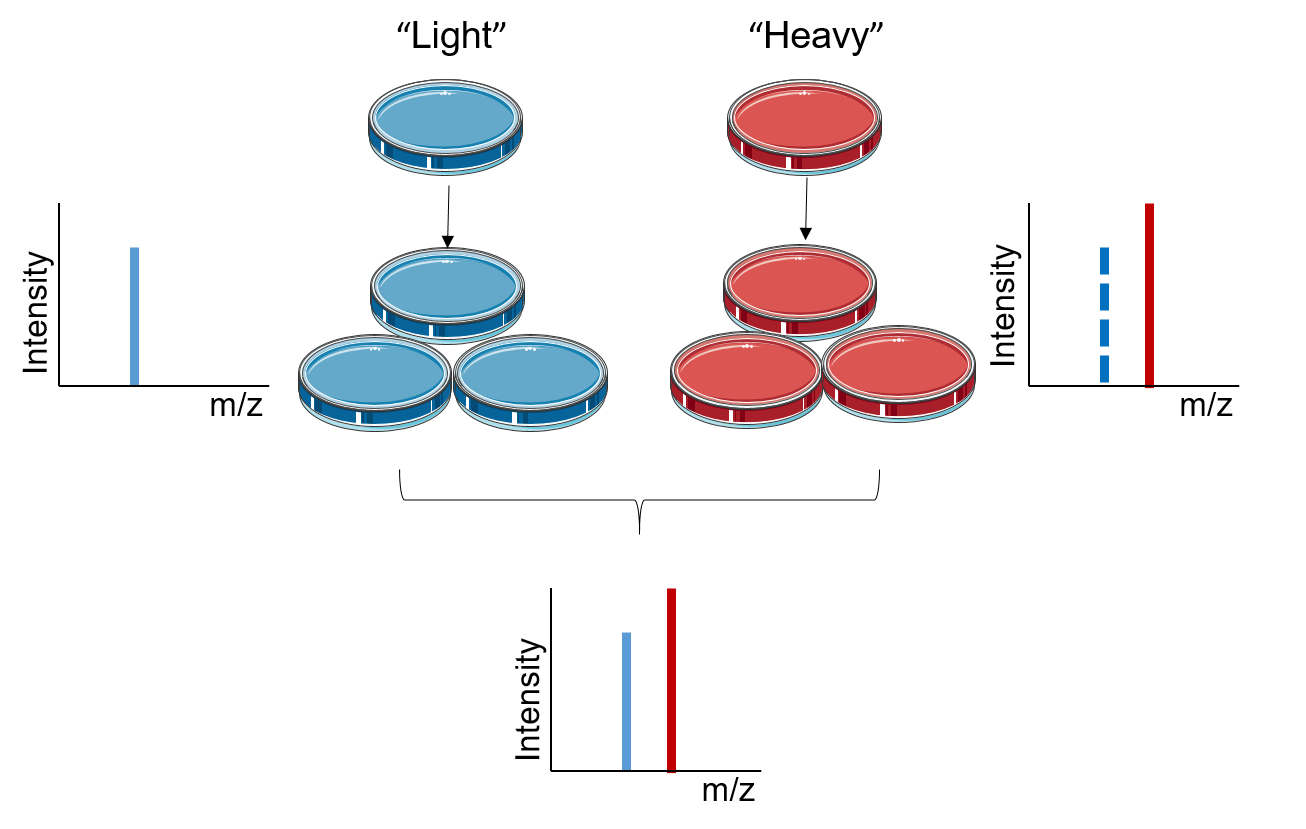
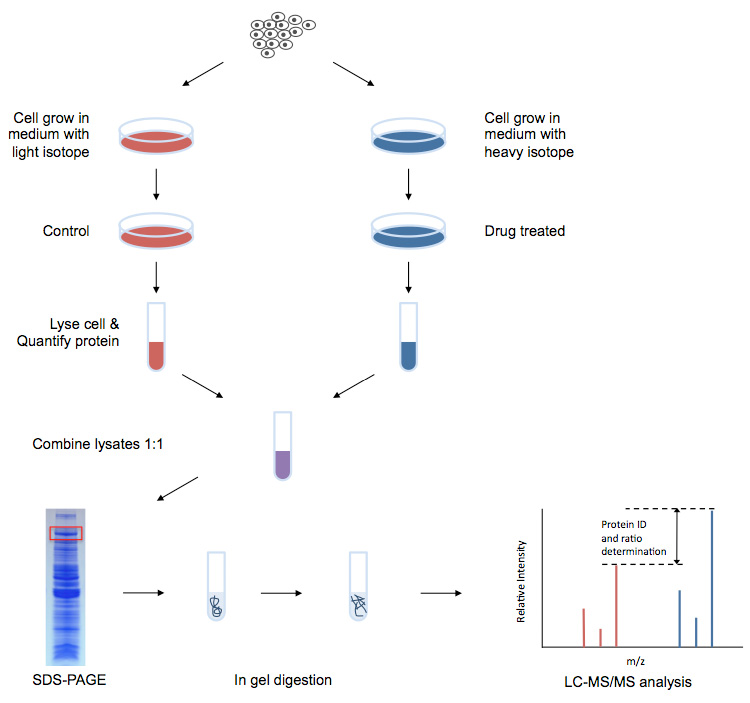
Stable Isotope Labeling Using Amino Acids in Cell Culture (SILAC): Principles, Workflow, and Applications – Creative Proteomics Blog

Prevention of Amino Acid Conversion in SILAC Experiments with Embryonic Stem Cells - Molecular & Cellular Proteomics

JCI Insight - Deep phenotyping of human induced pluripotent stem cell–derived atrial and ventricular cardiomyocytes

Determination of an Angiotensin II-regulated Proteome in Primary Human Kidney Cells by Stable Isotope Labeling of Amino Acids in Cell Culture ( SILAC) - Journal of Biological Chemistry

A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC) | Semantic Scholar

Amino acid based labeling techniques. (A) SILAC labeling. Proteins are... | Download Scientific Diagram

Quantitative proteomics using SILAC: Principles, applications, and developments - Chen - 2015 - PROTEOMICS - Wiley Online Library

IJMS | Free Full-Text | SILAC-Based Quantification of TGFBR2-Regulated Protein Expression in Extracellular Vesicles of Microsatellite Unstable Colorectal Cancers | HTML

SILAC Mouse for Quantitative Proteomics Uncovers Kindlin-3 as an Essential Factor for Red Blood Cell Function: Cell

Stable Isotope Labeling with Amino Acids in Cell Culture (SILAC) for Studying Dynamics of Protein Abundance and Posttranslational Modifications

An Enhanced In Vivo Stable Isotope Labeling by Amino Acids in Cell Culture ( SILAC) Model for Quantification of Drug Metabolism Enzymes * - Molecular & Cellular Proteomics
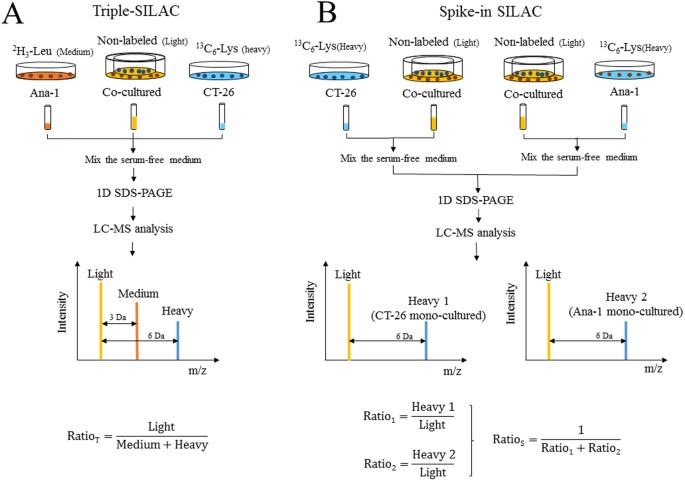
SILAC–based quantitative MS approach for real-time recording protein-mediated cell-cell interactions | Scientific Reports